Merging and interpolation of observations
Jens Daniel Müller
18 May, 2021
Last updated: 2021-05-18
Checks: 7 0
Knit directory: BloomSail/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20191021) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 00a2574. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: data/
Ignored: output/Plots/Figures_publication/.tmp.drivedownload/
Untracked files:
Untracked: data_os-2020-120_submission.zip
Unstaged changes:
Modified: code/Workflowr_project_managment.R
Modified: output/Plots/Figures_publication/Appendix/Fig_A1.pdf
Modified: output/Plots/Figures_publication/Appendix/Fig_A1.png
Modified: output/Plots/Figures_publication/Appendix/Fig_A2.pdf
Modified: output/Plots/Figures_publication/Appendix/Fig_B1.pdf
Modified: output/Plots/Figures_publication/Appendix/Fig_B1.png
Modified: output/Plots/Figures_publication/Appendix/Fig_B2.pdf
Modified: output/Plots/Figures_publication/Appendix/Fig_B2.png
Modified: output/Plots/Figures_publication/Appendix/Fig_C1.pdf
Modified: output/Plots/Figures_publication/Appendix/Fig_C2.pdf
Modified: output/Plots/Figures_publication/Appendix/Fig_C3.pdf
Modified: output/Plots/Figures_publication/Appendix/Fig_C3.png
Modified: output/Plots/Figures_publication/Appendix/Fig_C4.pdf
Modified: output/Plots/Figures_publication/Article/Fig_1.pdf
Modified: output/Plots/Figures_publication/Article/Fig_1.png
Modified: output/Plots/Figures_publication/Article/Fig_2.pdf
Modified: output/Plots/Figures_publication/Article/Fig_2.png
Modified: output/Plots/Figures_publication/Article/Fig_3.pdf
Modified: output/Plots/Figures_publication/Article/Fig_3.png
Modified: output/Plots/Figures_publication/Article/Fig_4.pdf
Modified: output/Plots/Figures_publication/Article/Fig_5.pdf
Modified: output/Plots/Figures_publication/Article/Fig_6.pdf
Modified: output/Plots/NCP_best_guess/tm_profiles_ID_pCO2_tem_sal_CT.pdf
Modified: output/Plots/NCP_best_guess/tm_profiles_pCO2_tem_sal_CT.pdf
Modified: output/Plots/response_time/all_plots.pdf
Modified: output/Plots/response_time/profiles_pCO2.pdf
Modified: output/Plots/response_time/profiles_pCO2_delta_grid.pdf
Modified: output/Plots/response_time/profiles_pCO2_delta_rel_grid.pdf
Modified: output/Plots/response_time/profiles_pCO2_grid.pdf
Modified: output/Plots/response_time/tau_determination_pCO2_corr_flushperiods_nls.pdf
Modified: output/Plots/response_time/time_series_depth_pCO2_corr_by_profile.pdf
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were made to the R Markdown (analysis/merging_interpolation.Rmd) and HTML (docs/merging_interpolation.html) files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | 00a2574 | jens-daniel-mueller | 2021-05-10 | Build site. |
| html | 61e452c | jens-daniel-mueller | 2021-04-16 | Build site. |
| html | 5f4fb9a | jens-daniel-mueller | 2021-02-20 | Build site. |
| html | 516b294 | jens-daniel-mueller | 2021-02-18 | Build site. |
| html | 70a8950 | jens-daniel-mueller | 2021-02-11 | Build site. |
| html | acba234 | jens-daniel-mueller | 2021-02-11 | Build site. |
| Rmd | 6d46720 | jens-daniel-mueller | 2021-02-11 | cleaning |
| html | d4ce04c | jens-daniel-mueller | 2021-02-11 | Build site. |
| Rmd | 0d87752 | jens-daniel-mueller | 2021-02-11 | cleaning |
| html | 4dfc4f8 | jens-daniel-mueller | 2021-02-11 | Build site. |
| Rmd | 8dc3356 | jens-daniel-mueller | 2021-02-11 | cleaning |
| html | 58c044c | jens-daniel-mueller | 2021-02-11 | Build site. |
| Rmd | fbe6ede | jens-daniel-mueller | 2021-02-11 | cleaning |
| html | 9544a3e | jens-daniel-mueller | 2021-02-11 | Build site. |
| Rmd | 775bef9 | jens-daniel-mueller | 2021-02-11 | cleaning |
| html | c0efe17 | jens-daniel-mueller | 2021-02-11 | Build site. |
| Rmd | 28f059f | jens-daniel-mueller | 2021-02-11 | cleaning |
| html | 1fe2dce | jens-daniel-mueller | 2021-02-11 | Build site. |
| Rmd | 4f7b259 | jens-daniel-mueller | 2021-02-11 | cleaning |
| html | 8e5e6d9 | jens-daniel-mueller | 2021-02-11 | Build site. |
| Rmd | 40f9a07 | jens-daniel-mueller | 2021-02-11 | cleaning |
| html | 8051798 | jens-daniel-mueller | 2021-02-10 | Build site. |
| html | c5fc34c | jens-daniel-mueller | 2021-01-22 | Build site. |
| html | 4277235 | jens-daniel-mueller | 2021-01-05 | Build site. |
| html | 9a3f42a | jens-daniel-mueller | 2020-10-24 | Build site. |
| html | 05248bf | jens-daniel-mueller | 2020-10-20 | Build site. |
| html | 1c4fe8e | jens-daniel-mueller | 2020-10-20 | table with time series in depth intervals added |
| html | 9e2ff85 | jens-daniel-mueller | 2020-10-13 | Build site. |
| Rmd | ff85457 | jens-daniel-mueller | 2020-10-13 | removed various Contros correction plots |
| html | 75d03b5 | jens-daniel-mueller | 2020-10-13 | Build site. |
| Rmd | 439d733 | jens-daniel-mueller | 2020-10-13 | removed various Contros correction plots |
| html | 6896725 | jens-daniel-mueller | 2020-10-01 | Build site. |
| html | 9f66019 | jens-daniel-mueller | 2020-10-01 | Build site. |
| html | 27c5431 | jens-daniel-mueller | 2020-09-29 | Build site. |
| Rmd | 2e0f902 | jens-daniel-mueller | 2020-09-29 | all parameters separate, rebuild |
| html | 1d01685 | jens-daniel-mueller | 2020-09-28 | Build site. |
| Rmd | d28129f | jens-daniel-mueller | 2020-09-28 | republish after tau factor set to 1 and using final pCO2 data |
| html | 4cfc1ad | jens-daniel-mueller | 2020-09-25 | Build site. |
| Rmd | 0e92cf9 | jens-daniel-mueller | 2020-09-25 | enabled chunks for merging and saving |
| html | 02a1609 | jens-daniel-mueller | 2020-09-25 | Build site. |
| Rmd | 99e69cf | jens-daniel-mueller | 2020-09-25 | activated read-in of th and ts data |
| html | 16554bc | jens-daniel-mueller | 2020-09-25 | Build site. |
| Rmd | 75e8c80 | jens-daniel-mueller | 2020-09-25 | plot with 10% sample size |
| html | 616c27f | jens-daniel-mueller | 2020-09-25 | updated repo manually |
| Rmd | 118f99e | jens-daniel-mueller | 2020-09-25 | comparison of pCO2 data included |
| html | 264566c | jens-daniel-mueller | 2020-09-25 | Build site. |
| Rmd | abc5bac | jens-daniel-mueller | 2020-09-25 | comparison of pCO2 data included |
| html | 904f0f7 | jens-daniel-mueller | 2020-09-23 | Build site. |
| Rmd | 7f497e4 | jens-daniel-mueller | 2020-09-23 | updated tau lm fit procedure |
| html | 8951791 | jens-daniel-mueller | 2020-09-23 | Build site. |
| Rmd | 9e87621 | jens-daniel-mueller | 2020-09-23 | included postprocessed cleaned data |
| html | ddd2d3e | jens-daniel-mueller | 2020-09-23 | Build site. |
| Rmd | 3ad71b0 | jens-daniel-mueller | 2020-09-23 | included postprocessed cleaned data |
| html | aea9be2 | jens-daniel-mueller | 2020-09-23 | Build site. |
| Rmd | ed17078 | jens-daniel-mueller | 2020-09-23 | included postprocessed cleaned data |
| html | 66bf52a | jens-daniel-mueller | 2020-09-23 | Build site. |
| Rmd | 0c8eed6 | jens-daniel-mueller | 2020-09-23 | included postprocessed cleaned data |
| html | c919fb7 | jens-daniel-mueller | 2020-06-29 | Build site. |
| Rmd | 1461cb6 | jens-daniel-mueller | 2020-06-29 | Fig update for talk |
| html | 603af23 | jens-daniel-mueller | 2020-05-25 | Build site. |
| html | 3414c23 | jens-daniel-mueller | 2020-05-25 | Build site. |
| html | 772e588 | jens-daniel-mueller | 2020-05-04 | Build site. |
| Rmd | 2ab39d7 | jens-daniel-mueller | 2020-05-04 | All profiles and timeseries in one plot pdf |
| html | 1ae50d3 | jens-daniel-mueller | 2020-05-04 | Build site. |
| Rmd | e78c435 | jens-daniel-mueller | 2020-05-04 | finalized time sync check |
| html | f95bf94 | jens-daniel-mueller | 2020-05-04 | Build site. |
| Rmd | 56f6c8a | jens-daniel-mueller | 2020-05-04 | corrected dep_maxgap removel criterion |
| html | c23350e | jens-daniel-mueller | 2020-05-04 | Build site. |
| Rmd | 3067532 | jens-daniel-mueller | 2020-05-04 | revise time sync |
| html | 3832733 | jens-daniel-mueller | 2020-04-30 | Build site. |
| Rmd | 4f4ab08 | jens-daniel-mueller | 2020-04-30 | harmonized code until RT determination |
| html | 6465570 | jens-daniel-mueller | 2020-04-29 | Build site. |
| Rmd | 0bbf0e6 | jens-daniel-mueller | 2020-04-29 | revised nomenclature |
| html | ebd1948 | jens-daniel-mueller | 2020-04-29 | Build site. |
| Rmd | 52090bf | jens-daniel-mueller | 2020-04-29 | correct interpolation, new d pco2 plot range |
| html | d9248a6 | jens-daniel-mueller | 2020-04-29 | Build site. |
| Rmd | 70bd3f0 | jens-daniel-mueller | 2020-04-29 | correct interpolation, new d pco2 plot |
| html | b5722a7 | jens-daniel-mueller | 2020-04-28 | Build site. |
| html | 472c2b4 | jens-daniel-mueller | 2020-04-21 | Build site. |
| html | f8fcf50 | jens-daniel-mueller | 2020-04-19 | created pub figures for time series |
| html | a6c4c22 | jens-daniel-mueller | 2020-03-30 | Build site. |
| html | 80c78b3 | jens-daniel-mueller | 2020-03-30 | Build site. |
| html | 5f8ca30 | jens-daniel-mueller | 2020-03-20 | Build site. |
| html | 2a20453 | jens-daniel-mueller | 2020-03-20 | Build site. |
| html | 473ab25 | jens-daniel-mueller | 2020-03-19 | Build site. |
| html | 81f022e | jens-daniel-mueller | 2020-03-18 | Build site. |
| html | 1e39d85 | jens-daniel-mueller | 2020-03-18 | Build site. |
| html | 2105236 | jens-daniel-mueller | 2020-03-18 | Build site. |
| html | 05b9bdc | jens-daniel-mueller | 2020-03-17 | Build site. |
| html | 0202742 | jens-daniel-mueller | 2020-03-16 | Build site. |
| html | 8e83afd | jens-daniel-mueller | 2020-03-12 | Build site. |
| html | a3ddea4 | jens-daniel-mueller | 2020-03-12 | Build site. |
| html | 52621ea | jens-daniel-mueller | 2020-03-12 | Build site. |
| html | e43a6f2 | jens-daniel-mueller | 2019-12-19 | Build site. |
| html | 3042ff3 | jens-daniel-mueller | 2019-12-19 | Build site. |
| Rmd | 282c3ac | jens-daniel-mueller | 2019-12-19 | whole data set RT corrected |
| html | 78710ee | jens-daniel-mueller | 2019-12-09 | Build site. |
| Rmd | c6cfca5 | jens-daniel-mueller | 2019-12-09 | RT correction incl OGB data |
| html | c6cfca5 | jens-daniel-mueller | 2019-12-09 | RT correction incl OGB data |
| html | bc6f19b | jens-daniel-mueller | 2019-11-22 | Build site. |
| Rmd | 03b1b97 | jens-daniel-mueller | 2019-11-22 | updated RT determination |
| html | 874dac5 | jens-daniel-mueller | 2019-11-22 | Build site. |
| Rmd | f875795 | jens-daniel-mueller | 2019-11-22 | now clean |
| html | d921065 | jens-daniel-mueller | 2019-11-14 | Build site. |
| Rmd | 252f84d | jens-daniel-mueller | 2019-11-14 | included EDA in data base |
| html | d61a468 | jens-daniel-mueller | 2019-11-14 | Build site. |
| html | f3277a5 | jens-daniel-mueller | 2019-11-08 | Build site. |
| html | 4256bcf | jens-daniel-mueller | 2019-11-08 | Build site. |
| html | 72687ee | jens-daniel-mueller | 2019-11-08 | Build site. |
| html | 74212a6 | jens-daniel-mueller | 2019-11-08 | Build site. |
| Rmd | 6cb1935 | jens-daniel-mueller | 2019-11-08 | response_time updated |
| html | 33e3659 | jens-daniel-mueller | 2019-10-22 | Build site. |
| Rmd | efcafd1 | jens-daniel-mueller | 2019-10-22 | Added data base, merging, and RT determination |
| html | 1595fe9 | jens-daniel-mueller | 2019-10-21 | Build site. |
| Rmd | 4131b9c | jens-daniel-mueller | 2019-10-21 | finisehd read CTD and HydroC, created merging Rmd |
library(tidyverse)
library(lubridate)
library(zoo)1 Scope of this script
- merge CTD, pCO2 and track data
- interpolate to common time stamp
- compare pCO2 from different source files
2 CTD (ts) + HydroC CO2 data (th, V1)
2.1 Merging summarized data sets
Before merging the ts and th data set, the time stamp of ts is adjusted to match exactly that of th, based on zeroing pCO2 values recorded in the analog output (ts) and the internal memory (th).
The th data set used here, is V1 of the post-processed data by KM Contros.
# Load Sensor and HydroC data ---------------------------------------------
ts <- read_csv(here::here("data/intermediate/_summarized_data_files",
"ts.csv"),
col_types = list("pCO2_analog" = col_double()))
th <- read_csv(here::here("data/intermediate/_summarized_data_files",
"th.csv"))
# Time offset correction ----------------------------------------------
# Time offset was determined by comparing zeroing reads from Sensor and th
# in the plots produced in the section Time stamp synchronicity below
# before applying this correction
ts <- ts %>%
mutate(day = yday(date_time),
date_time = if_else(day >= 206 & day <= 220,
date_time - 80, date_time - 10)) %>%
select(-day)
# Merge Sensor and HydroC data --------------------------------------------
ts_th <- full_join(ts, th) %>%
arrange(date_time)
rm(th, ts)A pdf with plots of the zeroing signals to check the time stamp syncronicity, can be found here:
2.2 Interpolation to common time stamp
CTD (ts) and auxillary recordings (15 sec measurment interval) are interpolated to the HydroC (th) time stamps (first 10 sec, than 1 sec measurement interval). Interpolation of ts data is not done when gaps between observations are larger than 20, indicating that th was running without ts, eg during data download from th. Thereafter, th readings not falling in regular transects/profilings are removed, by removing rows with NA depth values. Furthermore, ts readings without corresponding th readings are removed, except during periods when th was not operating.
# Interpolate Sensor data to HydroC time stamp
ts_th <- ts_th %>%
mutate(
dep_maxgap = na.approx(dep, na.rm = FALSE, maxgap = 20),
dep = approxfun(date_time, dep)(date_time),
sal = approxfun(date_time, sal)(date_time),
tem = approxfun(date_time, tem)(date_time),
pCO2_analog = approxfun(date_time, pCO2_analog)(date_time)
) %>%
#remove HC readings not falling in regular transects/profiling
filter(!is.na(dep_maxgap)) %>%
select(-dep_maxgap) %>%
fill(ID, type, station) %>%
# removes CTD readings without corresponding HydroC reading
filter(!is.na(deployment),!is.na(pCO2_analog))# Time stamp synchronicity
ts_th_Zero <- ts_th %>%
filter(Zero == 1 | Flush == 1 & duration < 120)
pdf(
file = here::here(
"output/Plots/merging_interpolation",
"Zero_time_synchronization.pdf"
),
onefile = TRUE,
width = 5,
height = 5
)
for (i_deployment in unique(ts_th$deployment)) {
#i_deployment <- unique(ts_th_Zero$deployment)[1]
ts_th_Zero_deployment <- ts_th_Zero %>%
filter(deployment == i_deployment)
for (i_Zero_counter in unique(ts_th_Zero_deployment$Zero_counter)) {
#i_Zero_counter <- unique(ts_th_Zero_deployment$Zero_counter)[1]
print(
ts_th_Zero_deployment %>%
filter(Zero_counter == i_Zero_counter) %>%
ggplot() +
geom_point(aes(date_time, pCO2_corr, col = "HydroC")) +
geom_point(aes(date_time, pCO2_analog, col = "analog")) +
labs(
title = paste("Depl: ", i_deployment,
" | Zero_counter: ", i_Zero_counter)
)
)
}
}
dev.off()
rm(ts_th_Zero,
ts_th_Zero_deployment,
i_deployment,
i_Zero_counter)2.3 Read cleaned, processed pCO2 data (V2)
A revised post-processed HydroC pCO2 data set was provided by KM Contros after applying a drift correction to the cleaned raw data, i.e. those without data recorded during configuration and testing of the sensor. This data set is referred to as V2. The post-processing was still based on pre- and post-deployment calibration results.
# Read Contros corrected data file, based on cleaned recordings and
# without water vapor correction
th_new_withoutAW_all <-
read_csv2(
here::here(
"data/input/TinaV/Sensor/HydroC-pCO2/corrected_Contros",
"parameter&pCO2s(method 43)_new_withoutAW.txt"
),
col_names = c(
"date_time",
"Zero",
"Flush",
"p_NDIR",
"p_in",
"T_control",
"T_gas",
"%rH_gas",
"Signal_raw",
"Signal_ref",
"T_sensor",
"pCO2_corr",
"Runtime",
"nr.ave"
)
) %>%
mutate(
date_time = dmy_hms(date_time),
Flush = as.factor(as.character(Flush)),
Zero = as.factor(as.character(Zero))
)
# slive every 10th data point to reduce number for plotting
th_new_withoutAW <- th_new_withoutAW_all %>%
slice(seq(1, n(), 10))
# load analog pCO2 data (raw)
th_pre_cleaning <-
read_csv(here::here(
"data/intermediate/_summarized_data_files",
"th_pre_cleaning.csv"
))
# slive every 10th data point to reduce number for plotting
th_pre_cleaning <- th_pre_cleaning %>%
slice(seq(1, n(), 10))
# slive every 10th data point to reduce number for plotting
ts_th_sub <- ts_th %>%
slice(seq(1, n(), 10))2.3.1 Comparison of pCO2 time series
2.3.1.1 Analog vs internal
Below, pCO2 time series for the analog output and the post-processed (ie drift corrected) data V1 are shown.
ggplot() +
geom_path(data = ts_th_sub,
aes(date_time, pCO2_corr, col = "HydroC V1")) +
geom_path(data = ts_th_sub,
aes(date_time, pCO2_analog, col = "analog CTD")) +
scale_color_brewer(palette = "Set1", name = "pCO2 record") +
coord_cartesian(ylim = c(0, 600)) +
labs(y = expression(pCO[2] ~ (µatm)), x = "") +
facet_wrap(~ deployment, scales = "free_x", ncol = 1)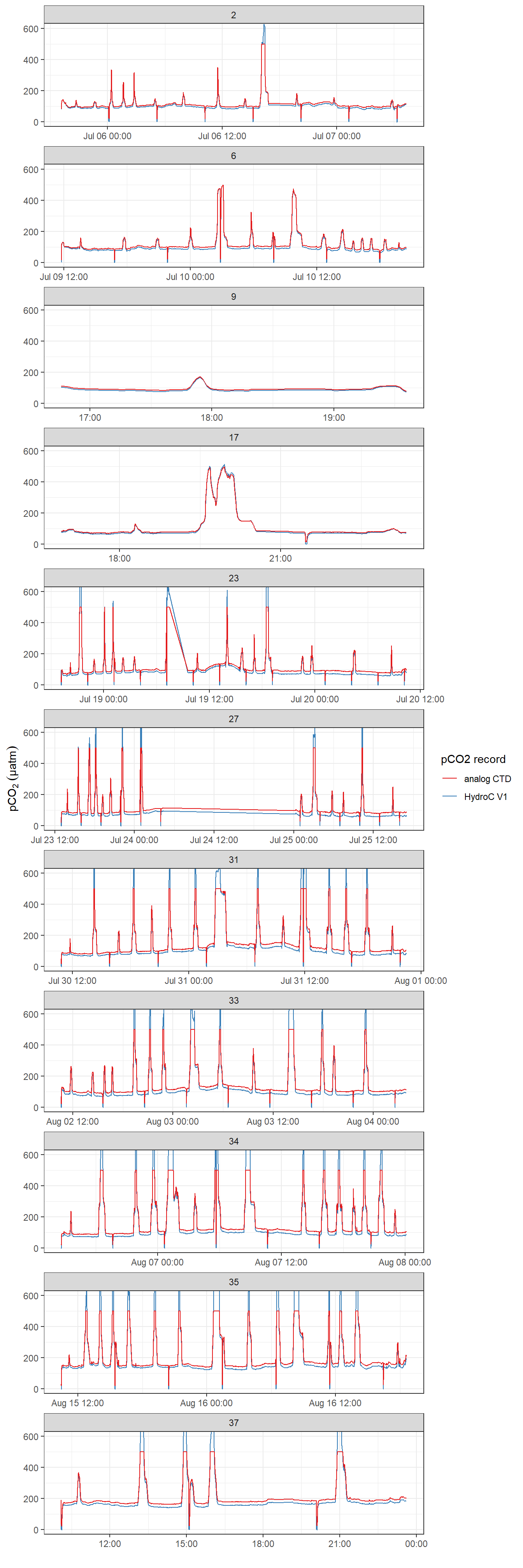
pCO2 record after interpolation to HydroC timestamp (analog output from HydroC recorded with CTD and drift corrected HydroC data V1 provided by Contos). ID refers to the starting date of each cruise. Please note that measurement range of the analog signal is technically restricted to 100-500 µatm. Zeroing periods are included.
2.3.1.2 Internal V1 vs. V2
Below, pCO2 time series for the post-processed (ie drift corrected) data V1 and V2 are shown.
th_comparison <- full_join(
ts_th_sub %>% select(date_time, deployment, pCO2_corr),
th_new_withAW %>% select(date_time, pCO2_corr) %>% rename(pCO2_withAW = pCO2_corr)
)
th_comparison <- full_join(
th_comparison,
th_new_withoutAW %>% select(date_time, pCO2_corr) %>% rename(pCO2_withoutAW = pCO2_corr)
)
th_comparison %>%
ggplot() +
geom_path(aes(date_time, pCO2_corr, col = "V1")) +
geom_path(aes(date_time, pCO2_withoutAW, col = "V2")) +
scale_color_brewer(palette = "Set1", name = "HydroC pCO2 (th)") +
coord_cartesian(ylim = c(0, 600)) +
labs(y = expression(pCO[2] ~ (µatm)), x = "") +
facet_wrap( ~ deployment, scales = "free_x", ncol = 1)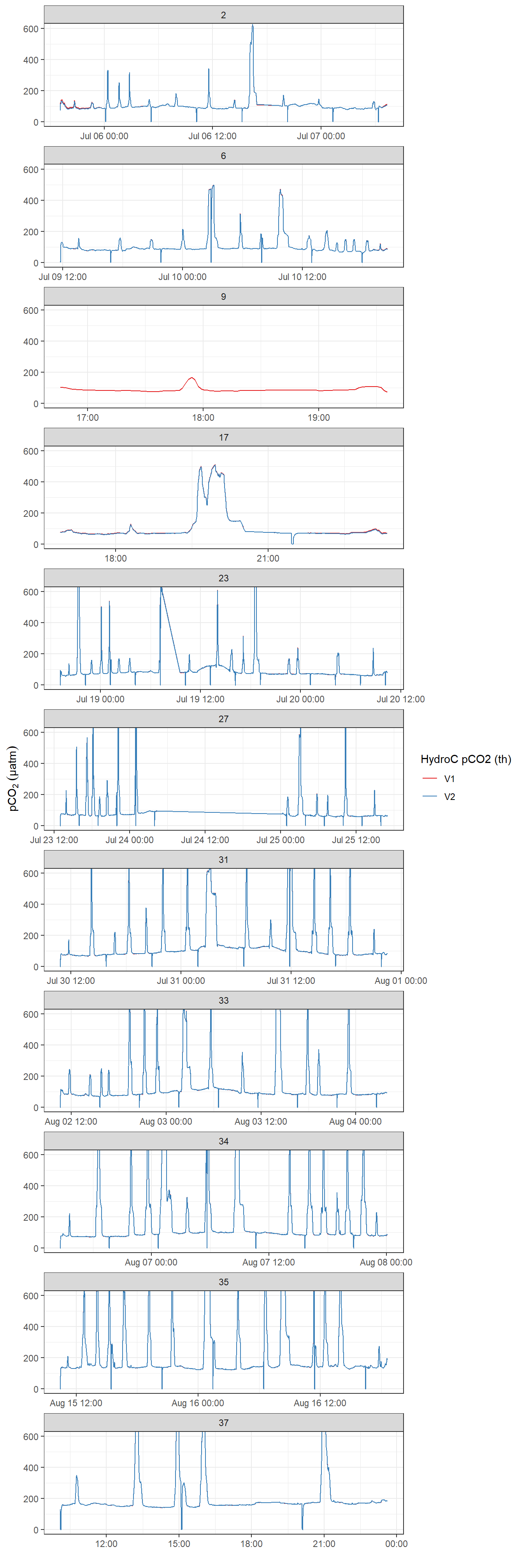
2.3.2 Replace pCO2 data
The pCO2 data V1 are replaced with V2, in the data set that will be used for further analysis.
th_new_withoutAW_all <- th_new_withoutAW_all %>%
select(date_time, pCO2_corr)
ts_th <- ts_th %>%
select(-pCO2_corr)
ts_th <- full_join(ts_th, th_new_withoutAW_all)
rm(th_new_withoutAW_all)2.3.3 Offset analog vs post-processed pCO2 (V2)
Below, the offset pCO2 time series between the analog output and the post-processed (ie drift corrected) data V2 are shown, which allows to judge the magnitude of the drift correction applied to V2.
ts_th %>%
ggplot() +
geom_path(aes(date_time, pCO2_corr - pCO2_analog)) +
ylim(-30, 0) +
labs(y = expression(pCO[2] ~ (µats_th)), x = "") +
facet_wrap( ~ deployment, scales = "free_x", ncol = 1)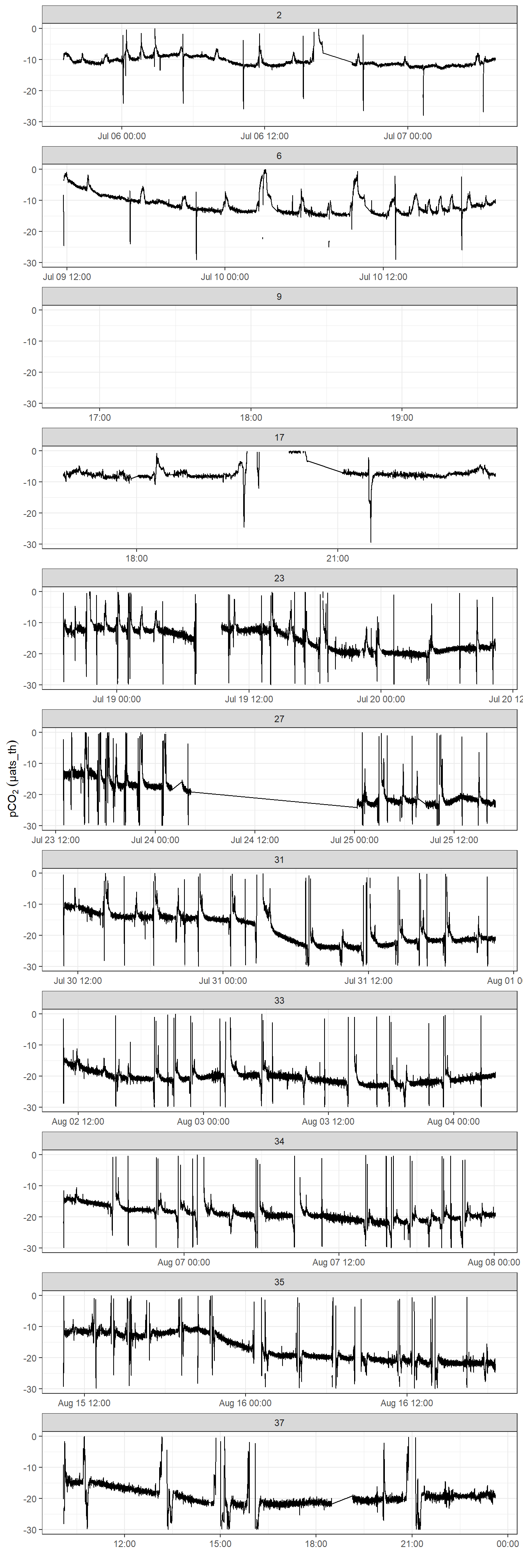
pCO2 difference between analog and drift corrected data provided by Contos. Please note that pCO2 range is restricted to 0 - 30 µatm.
3 Merge sensor (ts_th) + track (tt) data
tt <- read_csv(here::here("data/intermediate/_summarized_data_files",
"tt.csv"))
tm <- full_join(ts_th, tt) %>%
arrange(date_time)
# interpolate tt data and than remove columns that originate from tt time stamp
tm <- tm %>%
mutate(lat = approxfun(date_time, lat)(date_time),
lon = approxfun(date_time, lon)(date_time)) %>%
filter(!is.na(dep))4 Write merged file
tm %>% write_csv(here::here("data/intermediate/_merged_data_files/merging_interpolation",
"tm.csv"))
rm(tm, ts_th, tt)
sessionInfo()R version 4.0.3 (2020-10-10)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 19042)
Matrix products: default
locale:
[1] LC_COLLATE=English_Germany.1252 LC_CTYPE=English_Germany.1252
[3] LC_MONETARY=English_Germany.1252 LC_NUMERIC=C
[5] LC_TIME=English_Germany.1252
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] zoo_1.8-8 lubridate_1.7.9.2 forcats_0.5.0 stringr_1.4.0
[5] dplyr_1.0.2 purrr_0.3.4 readr_1.4.0 tidyr_1.1.2
[9] tibble_3.0.4 ggplot2_3.3.3 tidyverse_1.3.0 workflowr_1.6.2
loaded via a namespace (and not attached):
[1] Rcpp_1.0.5 here_1.0.1 lattice_0.20-41 ps_1.5.0
[5] assertthat_0.2.1 rprojroot_2.0.2 digest_0.6.27 R6_2.5.0
[9] cellranger_1.1.0 backports_1.2.1 reprex_0.3.0 evaluate_0.14
[13] highr_0.8 httr_1.4.2 pillar_1.4.7 rlang_0.4.10
[17] readxl_1.3.1 rstudioapi_0.13 whisker_0.4 rmarkdown_2.6
[21] labeling_0.4.2 munsell_0.5.0 broom_0.7.5 compiler_4.0.3
[25] httpuv_1.5.4 modelr_0.1.8 xfun_0.19 pkgconfig_2.0.3
[29] htmltools_0.5.0 tidyselect_1.1.0 fansi_0.4.1 crayon_1.3.4
[33] dbplyr_2.0.0 withr_2.3.0 later_1.1.0.1 grid_4.0.3
[37] jsonlite_1.7.2 gtable_0.3.0 lifecycle_0.2.0 DBI_1.1.0
[41] git2r_0.27.1 magrittr_2.0.1 scales_1.1.1 cli_2.2.0
[45] stringi_1.5.3 farver_2.0.3 fs_1.5.0 promises_1.1.1
[49] xml2_1.3.2 ellipsis_0.3.1 generics_0.1.0 vctrs_0.3.6
[53] RColorBrewer_1.1-2 tools_4.0.3 glue_1.4.2 hms_0.5.3
[57] yaml_2.2.1 colorspace_2.0-0 rvest_0.3.6 knitr_1.30
[61] haven_2.3.1